Biological Science Faculty Member
Dr. Hongchang Cui
- Office: 2020 King Life Sciences
- Office: (850) 645-1967
- Area: Cell and Molecular Biology
- Lab: King Life Sciences
- Lab: 2016 King Life Sciences Building
- Fax: (850) 645-8447
- Mail code: 4295
- E-mail: hcui@bio.fsu.edu

Associate Professor
Ph.D., Penn State University, 2003;
Postdoc, Duke University, 2009
Graduate Faculty Status
Research and Professional Interests:
- Research interests
Cell fate specification and reprogramming in plants; evolutionary and developmental biology; plant-environment interaction - Approaches
Genomics; epigenomics; proteomics; fluorescence-assisted cell sorting (FACS); molecular genetics - Model systems
Arabidopsis thaliana; Physcomitrella patens
1. Cell fate specification and reprogramming
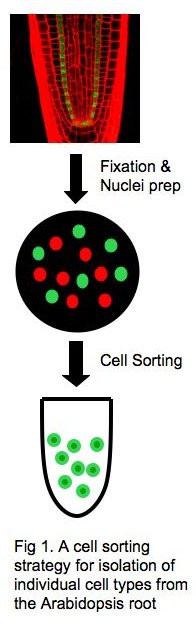
A major goal of my research is to elucidate the molecular basis of cell fate specification and reprogramming in plants. To this end, I will determine the global landscape of histone marks in individual cell types in the Arabidopsis root, using the ChIP/chip or ChIP-seq method. The root of Arabidopsis thaliana is best suited to these studies, as it permits isolation of individual cell types by fluorescence-assisted cell sorting (FACS) coupled with cell-type–specific GFP marker lines (1) (Fig. 1). Genes that are marked differently in different cell types will be identified, and their role in cell-fate specification will then be characterized by standard molecular-genetics methods.
Regenerative ability is an ancient trait, but somehow it has been lost in the majority of higher organisms, including humans. Higher plants are an exception, however: a whole plant can be regenerated from differentiated cells. Plant cells are thus believed to be totipotent. What makes plants so special, then? An important clue to this enigma came from a recent study, which showed that in animal stem cells the promoters of cell-fate-determination genes carry both active and repressive histone marks (termed “bivalent marks”), but in differentiated cell types only one of the marks is retained (2). This finding provides a plausible explanation why stem cells are developmentally plastic. It also suggests that totipotency genes in plants may have bivalent marks not only in stem cells but also in differentiated cells. With the cellular-epigenomics approach, we are now in an excellent position to address this important question.
2. Evolutionary and developmental biology
Mosses are among the most primitive land plants; they lack some of the critical features of other land plants such as roots and vascular tissues. Intriguingly, recent sequencing of the Physcomitrella patens genome revealed that this moss has clear homologs to the majority of Arabidopsis genes, including those involved in root development (3). The question is, what is the function of the root-development-related genes in the moss? I will address this question using P. patens as a model system, not only because its genome sequence is now available but also because it is amenable to genetic manipulation. As a matter of fact, P. patens is so far the only plant species with which the homology-based gene-targeting method can be efficiently performed, so it is a very powerful system for molecular evolutionary studies. By determining the function of the moss homologs of root development genes, we would gain significant insight into the origin of root. Physcomitrella will also be used as a model system for characterizing the genes that control totipotency, as generating transgenic plants from a single somatic cell is already routine.
3. Plant-environment interaction
Plants are sessile, and thus, to survive a changing environment, they must be able to adjust their developmental plan accordingly, in addition to mounting a whole array of stress responses. After a short exposure to a mild stress, such as cold, plants also acquire the ability to withstand a higher intensity of the same stress, a phenomenon called acclimation. Evidence indicates that plant hormones play an important role in this process, and one critical component of the regulatory mechanism is epigenetic regulation. Another goal of my research is therefore to reveal how plants integrate various environmental cues into their development program and the molecular basis of adaptation, with a focus on phytohormone signaling and epigenetic regulation.
1. K. Birnbaum et al., Nature Methods 2, 615. 2005.
2. B. E. Bernstein et al., Cell 125, 315. 2006.
3. S. A. Rensing et al., Science 319, 64. 2008.
Selected Publications:
Ma, F., Fu, J., Yang, T., Yang, L., Hu, X., Cui, H. (2025) Systematic analysis of the GRAS family genes in Arabidopsis thaliana reveals a role for SCL14 in primary root growth and redox homeostasis. Plant Journal. 124: e70534. DOI: 10.1111/tpj.70534.
Yang, T., Shi, X., Ma, F., Zhang, X., Li, S., Fu, J., Wu, S., Cui, H. (2025). OsIDD6, an INDETERMINATE DOMAIN containing transcription factor in rice, plays an essential role in reproductive development. Plant Journal. 123: e70408. DOI: 10.1111/tpj.70408.
Cui, H. (2025) Tender But Tough: How light and developmental stage cooperate to protect young leaves from the cold. Plant Cell and Environment. 48: 6962–6964. DOI: 10.1111/pce.15677.
Shrestha, R., Reyes, A.V., Carey, S., Karunadasa, S.S., Zhai, W., Byun, D., Lin, W.-D., Li, J., Alert, K., Cui, H., Wang, Z.-Y., Xu, S.-L. (2025) Next-Generation Mapping of the ACINUS-Mediated Alternative Splicing Machinery and Its Regulation by O-glycosylation in Arabidopsis. bioRxiv. DOI:10.1101/2025.01.04.63132.
Zhou, C., Yang, T., Cai, M., Cui, H., Yu, F., Liu, H., Fu, J. (2025) Comprehensive analysis of the INDETERMINATE DOMAIN (IDD) gene family in Marchantia polymorpha brings new insight into evolutionary developmental biology. BMC Genomics. 26: 415. DOI:10.1186/s12864-025-11609-7.
Zhu, X., Liu, L., Yang, L., Ma, F., Fu, J., Cui, H. (2025) ELONGATED HYPOCOTYL 5 promotes root growth by maintaining redox homeostasis and repressing oxidative stress response. Plant Physiology, 19(2): kiaf036. DOI:10.1093/plphys/kiaf036.
Shi, X., Yang, T., Ren, M., Fu, J., Bai, J., Cui, H. (2024) AT-hook motif nuclear localized transcription factors function redundantly in promoting root growth through modulation of redox homeostasis. Plant Journal. 120 (1): 199-217. DOI:10.1111/tpj.16981.
Fu, J., Zhou, C., Ma, F., Zhao, J., Yu, F., Cui, H. (2024) The PLETHORA homolog in Marchantia polymorpha is essential to meristem maintenance, developmental progression, and redox homeostasis. Plant Cell Physiology, 65 (8): 1231-1244. DOI:10.1093/pcp/pcae055 (Editor's choice & Featured on the cover).
Yang, L., Hu, X., Ren, M., Ma, F., Fu, J., Cui, H. (2024) Stem-cell expressed DEVIL-like small peptides maintain root growth under abiotic stress via ABA signaling. Plant Physiology. 194: 2372–2386. DOI:10.1093/plphys/kiad659.
Cui, H. (2023) NAC1 goes TOPLESS to manage context proliferation. Molecular Plant 16(5): 804-805.
DOI: 10.1016/j.molp.2023.04.001.
Wang, B., X. Shi, J. Gao, R. Liao, J. Fu, J. Bai, and H. Cui (2023) SCARECROW maintains the stem cell niche in Arabidopsis root by ensuring telomere integrity. Plant Physiology 192: 1115–1131. DOI:10.1093/plphys/kiad181 (On the cover).
Ishikawa, M., A. Fujiwara, K. Kosetsu, Y. Horiuchi , N. Kamamoto, N. Umakawa, Y. Tamadaa, L. Zhang, K. Matsushita, G. Gergo Palfalvi, T. Nishiyama, S. Kitasaki, Y. Masuda, Y. Shiroz, M. Kitagawa, H. Cui, S. Yoshida, Y. Hiwatashi, Y. Kabeya, S. Shigenobu, T. Aoyama, K. Kato, T. Murat, K. Fujimoto, P. N. Benfey, M. Hasebe, and R. Kofuji (2023) GRAS transcription factors regulate cell division planes in moss overriding the default rule. Proc. Natl. Acad. Sci 120: e22106321.DOI: 10.1073/pnas.2210632120
Cui, H., D. Kong, and Y. Hao (2023) Materials and methods for controlling bundle sheath cell fate and function in plants. US Patent No. 11542517.
Hu, X., L. Yang, M. Ren, L. Liu, J. Fu, and H. Cui (2022) TGA factors promote plant root growth by modulating redox homeostasis or response. Journal of Integrative Plant Biology 64: 1543-1559. DOI: 10.1111/jipb.13310.
Li, W., M. Zhang, L. Qiao, Y. Chen, D. Zhang, X. Jing, P. Gan, Y. Huang, J. Gao, W. Liu, C. Shi, H. Cui, H. Li, and K. Chen (2022) Characterization of wavy root 1, an agravitropism allele, reveals the functions of OsPIN2 in fine regulation of auxin transport and distribution and in ABA biosynthesis and response in rice (Oryza sativa L.). Crop Journal 10: 980-992. DOI: 10.1016/j.cj.2021.12.004.
Cui, H. (2021) Challenges and approaches to crop improvement through C3-to-C4 engineering. Frontiers in Plant Science 12: Article 715391. DOI: 10.3389/fpls.2021.715391.
Fu, J., X. Zhang, J. Liu, X. Gao, J. Bai, Y. Hao, and H. Cui (2021) A mechanism coordinating root elongation, endodermal differentiation, redox homeostasis and response. Plant Journal 107: 1029-1039. DOI:10.1111/tpj.15361.
Gao, X., J. Fu, and H. Cui (2021) Analysis of the expression pattern of some Liverwort MYB genes in different tissues at various developmental stages and their responses to plant hormone treatments. Acta Agriculturae Boreali-Occidentalis Sinica 30: 109-115. (Abstract)
Wang, B., X. Shi, J. Gao, R. Liao, J. Fu, J. Bai, and H. Cui (2021) SCARECROW maintains the stem cell niche in Arabidopsis root by ensuring telomere integrity. bioRxiv 10.1101/2021.10.25.465752v1. DOI: 10.1101/2021.10.25.465752.
Cui, H., D. Kong, and Y. Hao (2020) Materials and methods for controlling bundle sheath cell fate and function in plants.. US Patent No. 10865420.
(Link)
Li, J., B. Wang, X. Zhu, R. Li, J. Fu, and H. Cui (2020) Novel regulators of sugar-mediated lateral root development in Arabidopsis thaliana. Genes 11: 1257. DOI:10.3390/genes11111257.
Bao, Z., J. Bai, H. Cui, and C. Gong (2019) A missing link in radial ion transport: ion transporters in the endodermis. Frontiers in Plant Science 10: 1-8. DOI: 10.3389/fpls.2019.00713.
Cui, H. (2016) Middle cortex formation in the root: an emergin picture of integrated regulatory mechanisms. Molecular Plant 9: 771-773. DOI: 10.1016/j.molp.2016.05.002.
Kong, D., Y. Hao, and H. Cui (2016) The WUSCHEL Related Homeobox Protein WOX7 Regulates the Sugar Response of Lateral Root Development in Arabidopsis thaliana. Molecular Plant 9: 261-270. DOI:https://doi.org/10.1016/j.molp.2015.11.006.
Cui, H. (2015) Cortex proliferation in plant root is a protective mechanism against abiotic stress in soil. Plant Signaling & Behavior 10: e1011949. DOI:10.1080/15592324.2015.1011949.
Cui, H., D. Kong, X. Liu, and Y. Hao (2014) SCARECROW, SCR-LIKE 23 and SHORT-ROOT control bundle sheath cell fate and function in Arabidopsis thaliana. Plant Journal 78: 319-327. DOI: 10.1111/tpj.12470.
Cui, H., D. Kong, P. Wei, Y. Hao, K. U. Torii, J. Lee, and J. Li (2014) SPINDLY, ERECTA and its ligand STOMAGEN have a role in redox-mediated cortex proliferation in the Arabidopsis root. Molecular Plant 7: 1727-1739. DOI: 10.1093/mp/ssu106 (on the cover).
Gao, X., C. Wang, and H. Cui (2014) Identification of bundle sheath cell-fate factors provides new tools for C3-to-C4 engineering. Plant Signaling & Behavior 9: e29162. DOI: 10.4161/psb.29162.
Cui, H., Y. Hao, and D. Kong (2012) SCARECROW has a SHORT-ROOT independent role in modulating the sugar response. Plant Physiology 158: 1769-1788. DOI: 10.1104/pp.111.191502.
Cui, H. (2012) Killing two birds with one stone - Transcriptional regulators coordinate development and stress responses in plants. Plant Signaling & Behavior 7: 701-703. DOI: 10.4161/psb.20283.
Hao, Y., and H. Cui (2012) SHORT-ROOT regulates vascular patterning, but not apical meristematic activity, in the Arabidopsis root through cytokinin homeostasis. Plant Signaling & Behavior 7: 314-317. DOI: 10.4161/psb.19118.
Cui, H. 2011 The Epigenetic Basis of Cell-Fate Specification and Reprogramming. In Epigenetics: A Reference Manual. Craig, J.M and Wong N.C. Caister Academic Press. Pp182-194.
(Abstract)
Cui, H., Y. Hao, M. Kovtun, V. Stolc, X. Deng, H. Sakakibara, and M. Kojima (2011) Genomewide direct target analysis reveals a role for SHORT-ROOT in root vascular patterning through cytokinin homeostasis. Plant Physiology 157: 1221-1231. DOI: 10.1104/pp.111.183178 (on the cover) (Selected as a recommended read by Faculty of 1000).
Iyer-Pascuzzi, A. S., T. Jackson, H. Cui, J. J. Petricka, W. Busch, H. Tsukagoshi, and P. N. Benfey (2011) Cell identity regulators link development and stress responses in the Arabidopsis root. Developmental Cell 21: 770-782. DOI: 10.1016/j.devcel.2011.09.009.
Sozzani, R., H. Cui (co-first author), M. A. Moreno-Risueno, W. Busch, J. M. Van Norman, T. Vernoux, S. M. Brady, W. Dewitte, J. A. Murray, and P. N. Benfey (2010) Spatiotemporal regulation of cell-cycle genes by SHORT-ROOT links patterning and growth. Nature 466: 128-132. DOI: 10.1038/nature09143.
Cui, H., and P. N. Benfey (2009) Interplay between SCARECROW, GA and LIKE HETEROCHROMATIN PROTEIN 1 in ground tissue patterning in the Arabidopsis root. Plant Journal 58: 1016-1027. DOI: 10.1111/j.1365-313X.2009.03839.x.
Cui, H., and P. N. Benfey (2009) Cortex proliferation: simple phenotype, complex regulatory mechanisms. Plant Signaling & Behavior 4: 551-553. DOI: 10.1111/j.1365-313X.2009.03839.x.
Cui, H., M. Levesque, T. Vernoux, J. Y. Wang, I. Blilou, B. Scheres, and P. N. Benfey (2007) An evolutionarily conserved mechanism delimiting SHR movement defines a single layer of endodermis in plants. Science 316: 421-425 (Article highlighted in Nature, Science, Cell, JCB and etc). DOI: 10.1126/science.1139531) (Selected as a must-read by Faculty of 1000.
Levesque, M. P., T. Vernoux, W. Busch, H. Cui, J. Y. Wang, I. Blilou, H. Hassan, K. Nakajima, N. Matsumoto, J. U. Lohmann, B. Scheres, and P. N. Benfey (2006) Whole-genome analysis of the SHORT-ROOT developmental pathway in Arabidopsis. PLoS Biology 4: e143. DOI: 10.1371/journal.pbio.0040143 (Selected as a recommended read by Faculty of 1000).
Cui, H., and N. V. Fedoroff (2002) Inducible DNA demethylation mediated by the maize Spm transposon-encoded TnpA protein. Plant Cell 14: 2883-2899. DOI: 10.1105/tpc.006163.
Cui, H. (in press) C3-to-C4 engineering: the next wave of green revolution In Trivedi, P.K. and Nath, P., John Willey & Sons
