Biological Science Faculty Member - Retired
Dr. Peter Fraser
- Office: 3062 King Life Sciences
- Office: (850) 644-5037
- Area: Cell and Molecular Biology
- Lab: King Life Sciences
- Lab: (850) 644-9816
- Fax: (850) 645-8447
- Mail code: 4295
- E-mail: pfraser@bio.fsu.edu

Professor
PhD, Molecular Biology. University of Pennsylvania, 1988.
Research and Professional Interests:
Dynamic changes in chromatin and chromosome architecture regulates patterns of cellular gene expression during differentiation and development, or in response to environmental signals. Our research looks at various levels of chromatin, chromosome and nuclear structure, from individual nucleosome modifications to the dynamic 3D structure of chromosomes and their inter-relationships in the nucleus and how they affect genome functions.
Single-cell Hi-C
The spatial organization of the genome inside the cell nucleus is tissue-specific and has been linked to several nuclear processes including gene activation, gene silencing, genomic imprinting, gene co-regulation, genome maintenance, DNA replication, DNA repair, immunoglobulin gene rearrangement, chromosomal translocations and X chromosome inactivation. Most chromosome conformaiton capture methods meaure populations of millions of cells, and since genome organization is highly variable from cell to cell these methods report on average or probabalistic confromations. To overcome this we developed single-cell Hi-C to assess genome conformation in single cells. This powerful method has allowed us to begin to understand the variability in chromosome conformation and chromosome dynamics.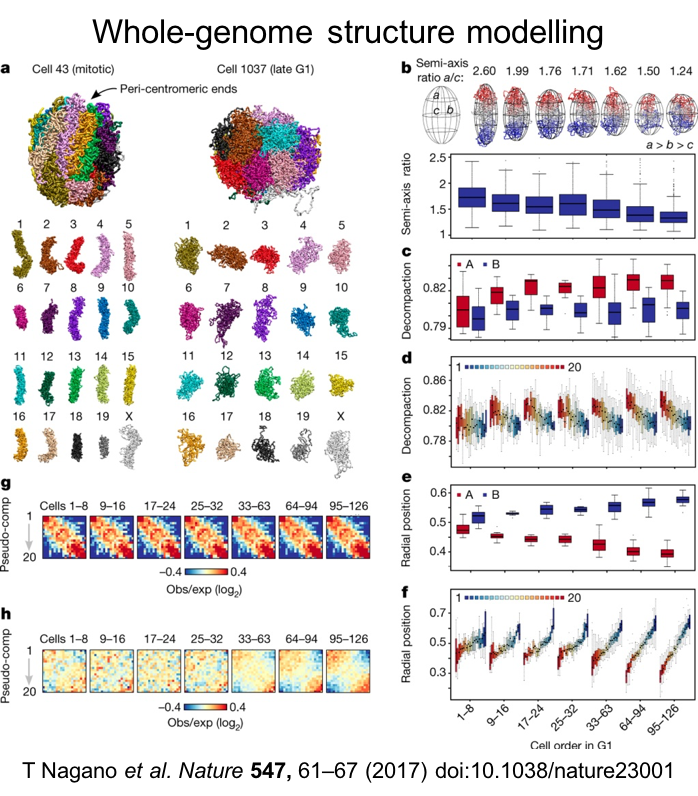
Promoter Capture Hi-C
Hi-C libraries are incredibly complex and cannot really identify chromatin interactions between distant genomic sequences at the restriction fragment level with statistical significance unless tens of billions or read-pairs are produced. To get around this shortcoming and to identify distal sequences that significantly interact with gene promoters using a statistical threshold we developed Promoter Capture Hi-C. This involves sequence capture using thousands of biotinylated RNA oligomers complementary to all annotated promoters to specifically enrich for promoter-containing fragments from Hi-C libraries. The enriched promoter-containing fragment libraries are then sequenced with high-throughput paired-end sequencing to reveal long-range, promoter-interacting genomic regions such as enhancers and other potential distal regulatory elements, as well as non-random, promoter-to-promoter contacts in the 3D nuclear space. We have used this data to create a greater understanding of global gene expression control, to assign distal regulatory elements to their target genes, to link human disease-associated variants to their target genes and to assess genome-wide contacts between promoters, which play roles in control of gene transcription and gene repression.
Full list of publications on Google Scholar
https://scholar.google.co.uk/citations?user=GuG93YUAAAAJ&hl=en
Selected Publications:
Choy MK, Javierre BM, Williams SG, Baross SL, Liu Y, Wingett SW, Akbarov A, Wallace C, Freire-Pritchett P, Rugg-Gunn PJ, Spivakov M, Fraser P, Keavney BD. (2018) Promoter interactome of human embryonic stem cell-derived cardiomyocytes connects GWAS regions to cardiac gene networks. Nat Commun. 2018 9:2526.
Beekman R, Chapaprieta V, Russiñol N, Vilarrasa-Blasi R, Verdaguer-Dot N, Martens JHA, Duran-Ferrer M, Kulis M, Serra F, Javierre BM, Wingett SW, Clot G, Queirós AC, Castellano G, Blanc J, Gut M, Merkel A, Heath S, Vlasova A, Ullrich S, Palumbo E, Enjuanes A, Martín-García D, Beà S, Pinyol M, Aymerich M, Royo R, Puiggros M, Torrents D, Datta A, Lowy E, Kostadima M, Roller M, Clarke L, Flicek P, Agirre X, Prosper F, Baumann T, Delgado J, López-Guillermo A, Fraser P, Yaspo ML, Guigó R, Siebert R, Martí-Renom MA, Puente XS, López-Otín C, Gut I, Stunnenberg HG, Campo E, Martin-Subero JI. (2018) The reference epigenome and regulatory chromatin landscape of chronic lymphocytic leukemia. Nat Med. 24:868-880.
Rivera-Mulia JC, Dimond A, Vera D, Trevilla-Garcia C, Sasaki T, Zimmerman J, Dupont C, Gribnau J, Fraser P, Gilbert DM. (2018) Allele-specific control of replication timing and genome organization during development. Genome Res. 28:800-811.
Schoenfelder S, Javierre BM, Furlan-Magaril M, Wingett SW, Fraser P. (2018) Promoter Capture Hi-C: High-resolution, Genome-wide Profiling of Promoter Interactions. J Vis Exp. Jun 28;(136). doi: 10.3791/57320.
Novo CL, BM Javierre, J Cairns, A Segonds-Pichon, SW Wingett, P Freire-Pritchett, M Furlan-Magaril, S Schoenfelder, P Fraser, P J Rugg-Gunn (2018) Long-Range Enhancer Interactions Are Prevalent in Mouse Embryonic Stem Cells and Are Reorganized upon Pluripotent State Transition. Cell Reports 22, 2615-2627.
Comoglio F, ParK HJ, Schoenfelder S, Barozzi I, Bode D, Fraser P, Green AR (2018) Thrombopoietin signaling to chromatin elicits rapid and pervasive epigenome remodeling within poised chromatin architectures. Genome Res 28, 295-309.
Wutz G, Várnai C, Nagasaka K, Cisneros DA, Stocsits RR, Tang W, Schoenfelder S, Jessberger G, Muhar M, Hossain MJ, Walther N, Koch B, Kueblbeck M, Ellenberg J, Zuber J, Fraser P, Peters J-M. (2017) Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 36,3573-3599.
Burren OS, Rubio García A, Javierre BM, Rainbow DB, Cairns J, Cooper NJ, Lambourne JJ, Schofield E, Castro Dopico X, Ferreira RC, Coulson R, Burden F, Rowlston SP, Downes K, Wingett SW, Frontini M, Ouwehand WH, Fraser P, Spivakov M, Todd JA, Wicker LS, Cutler AJ, Wallace C. (2017) Chromosome contacts in activated T cells identify autoimmune disease candidate genes. Genome Biol. 18:165
Rubin AJ, Barajas BC, Furlan-Magaril M, Lopez-Pajares V, Mumbach MR, Howard I, Kim DS, Boxer LD, Cairns J, Spivakov M, Wingett SW, Shi M, Zhao Z, Greenleaf WJ, Kundaje A, Snyder M, Chang HY, Fraser P, Khavari PA. (2017) Lineage-specific dynamic and pre-established enhancer-promoter contacts cooperate in terminal differentiation. Nat Genet. 49:1522-1528.
Nagano T, Lubling Y, Várnai C, Dudley C, Leung W, Baran Y, Mandelson-Cohen N, Wingett S, Fraser P and Tanay A. (2017) Cell cycle dynamics of chromosomal organisation at single-cell resolution Nature 547, 61-67.
Petersen R, Lambourne JJ, Javierre BM, Grassi L, Kreuzhuber R, Ruklisa D, Rosa IM, Tomé AR, Elding H, van Geffen JP, Jiang T, Farrow S, Cairns J, Al-Subaie AM, Ashford S, Attwood A, Batista J, Bouman H, Burden F, Choudry FA, Clarke L, Flicek P, Garner SF, Haimel M, Kempster C, Ladopoulos V, Lenaerts AS, Materek PM, McKinney H, Meacham S, Mead D, Nagy M, Penkett CJ, Rendon A, Seyres D, Sun B, Tuna S, van der Weide ME, Wingett SW, Martens JH, Stegle O, Richardson S, Vallier L, Roberts DJ, Freson K, Wernisch L, Stunnenberg HG, Danesh J, Fraser P, Soranzo N, Butterworth AS, Heemskerk JW, Turro E, Spivakov M, Ouwehand WH, Astle WJ, Downes K, Kostadima M, Frontini M. (2017) Platelet function is modified by common sequence variation in megakaryocyte super enhancers. Nat Commun. Jul 13;8:16058.
Harewood L, Kishore K, Eldridge MD, Wingett S, Pearson D, Schoenfelder S, Collins VP, Fraser P. (2017) Hi-C as a tool for precise detection and characterisation of chromosomal rearrangements and copy number variation in human tumours. Genome Biol. 18:125.
Siersbæk R, Madsen JGS, Javierre BM, Nielsen R, Bagge EK, Cairns J, Wingett SW, Traynor S, Spivakov M, Fraser P, Mandrup S. (2017) Dynamic Rewiring of Promoter-Anchored Chromatin Loops during Adipocyte Differentiation. Mol Cell 66,420-435.
Freire-Pritchett P, Schoenfelder S, Várnai C, Wingett SW, Cairns J, Collier AJ, García-Vílchez R, Furlan-Magaril M, Osborne CS, Fraser P, Rugg-Gunn P, Spivakov M. (2017) Global reorganisation of cis-regulatory units upon lineage commitment of human embryonic stem cells. eLife 6.
Aymard F, Aguirrebengoa M, Guillou E, Javierre BM, Bugler B, Arnould C, Rocher V, Iacovoni JS, Biernacka A, Skrzypczak M, Ginalski K, Rowicka M, Fraser P, and Legube G. (2017) Genome wide mapping of long range contacts unveils DNA Double Strand Breaks clustering at damaged active genes. Nat Struct & Mol Biol. doi:10.1038/nsmb.3387.
Javierre BM, Burren OS, Wilder SP, Kreuzhuber R, Hill SM, Sewitz S, Cairns J, Wingett SW, Várnai C, Thiecke MJ, Burden F, Farrow S, Cutler AJ, Rehnstrom K, Downes K, Grassi L, Kostadima M, Freire-Pritchett P, Wang F, The BLUEPRINT Consortium Stunnenberg HG, Todd JA, Zerbino DR, Stegle O, Ouwehand WH, Frontini M, Wallace C, Spivakov M and Fraser P. (2016) Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167:1369-1384.
Stunnenberg HG, The International Human Epigenome Consortium, and Hirst M. (2016) The International Human Epigenome Consortium: A Blueprint for Scientific Collaboration and Discovery. Cell 167:1145-1149.
Spivakov M and Fraser P (2016) Defining cell type with epigenetic profiling. Nature Biotech. 34:1126-1128.
Fraser P. (2016) Turning the tide on 3D nuclear organization. Nat Rev Mol Cell Biol. 17:738-738.
McGovern A, Schoenfelder S, Martin P, Massey J, Duffus K, Plant D, Yarwood A, Pratt A, Anderson A, Isaacs J, Diboll J, Thalayasingam N, Ospelt C, Barton A, Worthington J, Fraser P, Eyre S and Orozco G (2016) Capture Hi-C identifies a novel causal gene, IL20RA, in the pan-autoimmune genetic susceptibility region 6q23. Genome Biol. 17:212.
Cairns J, Freire-Pritchett P, Wingett SW, Dimond A, Plagnol V, Zerbino D, Schoenfelder S, Javierre BM, Osborne C, Fraser P, Spivakov M (2016) CHiCAGO: Robust Detection of DNA Looping Interactions in Capture Hi-C data, Genome Biol17:127.
Pancaldi V, de Santa Pau EC, Javierre BM, Juan D, Fraser P, Spivakov M, Valencia V, Rico D. (2016) Chromatin assortativity: Integrating epigenomic data and 3D genomic structure. Genome Biol 17:152.
Wilson NK, Schoenfelder S, Hannah R, Castillo MS, Schütte J, Ladopoulos V, Mitchelmore J, Goode DK, Calero-Nieto FJ, Moignard V, Wilkinson AC, Jimenez-Madrid I, Kinston S, Spivakov M, Fraser P, Göttgens B. (2016) Integrated genome-scale analysis of the transcriptional regulatory landscape in a blood stem/progenitor cell model. Blood DOI: http://dx.doi.org/10.1182/blood-2015-10-677393
Martin P, McGovern A, Orozco G, Duffus K, Yarwood A, Schoenfelder S, Cooper NJ, Barton A, Wallace C, Fraser P, Worthington J, Eyre S (2015) Capture Hi-C reveals novel candidate genes and complex long-range interactions with related autoimmune risk loci. Nature Commun 6:10069.
Nagano T, Lubling Y, Yaffe E, Wingett SW, Dean W, Tanay A and Fraser P (2015) Single-cell Hi-C for genome-wide detection of chromatin interactions that occur simultaneously in a single cell. Nature Protocols 10, 1986-2003.
Nagano T, Várnai C, Schoenfelder S, Javierre B-M, Wingett SW, Fraser P (2015) Comparison of Hi-C results using in-solution versus in-nucleus ligation. Genome Biol, 16:175.
Schoenfelder S, Sugar R, Dimond A, Javierre B-M, Armstrong H, Mifsud B, Dimitrova E, Matheson L, Tavares-Cadete F, Furlan-Magaril M, Segonds-Pichon A, Jurkowski W, Wingett SW, Tabbada K, Andrews S, Herman B, LeProust E, Osborne CS, Koseki H, Fraser P, Luscombe NM, Elderkin S. (2015) Polycomb repressive complex PRC1 spatially constrains the mouse embryonic stem cell genome. Nat Genet. 47, 1179-1186.
Mifsud B, Tavares-Cadete F, Young AN, Sugar R, Schoenfelder S, Ferreira L, Wingett SW, Andrews S, Grey W, Ewels PA, Herman B, Happe S, Higgs A, LeProust E, Follows GA, Fraser P, Luscombe NM, Osborne CS (2015) Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C. Nat Genet. 47, 598-606.
Furlan-Magaril M, Várnai C, Nagano T, Fraser P (2015) 3D genome architecture from populations to single cells. Curr Op in Genet & Dev 31, 36-41.
Schoenfelder S, Furlan-Magaril M, Mifsud B, Tavares-Cadete F, Sugar R, Javierre B-M, Nagano T, Katsman Y, Sakthidevi M, Wingett SW, Dimitrova E, Dimond A, Edelman LB, Elderkin S, Tabbada K, Darbo E, Andrews S, Herman B, Higgs A, LeProust E, Osborne CS, Mitchell JA, Luscombe NM, Fraser P (2015) The pluripotent regulatory circuitry connecting promoters to their long-range interacting elements. Genome Res 2015:gr.185272.114.
Jäger R, Migliorini G, Henrion M, Kandaswamy R, Speedy HE, Heindl A, Whiffin N, Carnicer MJ, Broome L, Dryden N, Nagano T, Schoenfelder S, Enge M, Yuan Y, Taipale J, Fraser P, Fletcher O, Houlston RS (2015) Capture Hi-C (cHi-C) identifies the chromatin interactome of colorectal cancer risk loci. Nat Commun 6:6178.
Chandra T, Ewels PA, Schoenfelder S, Furlan-Magaril M, Wingett SW, Kirschner K, Thuret J-Y, Andrews S, Fraser P, Reik W. (2015) Global Reorganization of the Nuclear Landscape in Senescent Cells. Cell Reports 10 (4), 471-483.
Dryden NH, Broome LR, Dudbridge F, Johnson N, Orr N, Schoenfelder S, Nagano T, Andrews S, Wingett S, Kozarewa I, Assiotis I, Fenwick K, Maguire SL, Campbell J, Natrajan R, Lambros M, Perrakis E, Ashworth A, Fraser P, and Fletcher O. (2014) Unbiased analysis of potential targets of breast cancer susceptibility loci by Capture Hi-C. Genome Res. 24: 1854-1868.
Nagano T, Lubling Y, Stevens TJ, Schoenfelder S, Yaffe S, Dean W, Laue ED, Tanay A and Fraser P. (2013) Single cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64.
A Dimond, and P Fraser (2013) Long Noncoding RNAs Xist in Three Dimensions. Science 341: 720-721.
Sexton T, Kurukuti S, Mitchell JA, Umlauf D, Nagano T and Fraser P. (2012) Sensitive detection of chromatin coassociations using enhanced chromosome conformation capture on chip. Nat Protoc 7:1335-1350.
Davison LJ, Wallace C, Cooper JD, Cope NF, Wilson NK, Smyth DJ, Howson JMM, Saleh N, Al-Jeffery A, Angus KL, Stevens HE, Nutland S, Duley S, Coulson RMR, Walker NM, Burren OS, Rice CM, Cambien F, Zeller T, Munzel T, Lackner K, Blankenberg S Gutenberg Heart Study, The Cardiogenics Consortium, Fraser P, Gottgens B and Todd JA. (2012) Long-range DNA looping and gene expression analyses identify DEXI as an autoimmune disease candidate gene. Hum Mol Genet. 21:322-333.
Eskiw C and Fraser P. (2011) Ultra-structural study of transcription factories in mouse erythroblasts. J Cell Sci. 124: 3676-3683.
Kang J, Xu B, Yao Y, Lin W, Hennessy C, Fraser P and Feng J. (2011) A Dynamical Model Reveals Gene Co-localizations in Nucleus. PLoS Comp Biol 7(7): e1002094.
Nagano T, and Fraser P (2011) No-nonsense functions for long non-coding RNAs. Cell, 145:178-181.
Harrigan JA, Belotserkovskaya R, Coates J, Dimitrova DS, Polo SE, Bradshaw CR, Fraser P, and Jackson SP (2011) Replication stress induces 53BP1-containing OPT domains in G1 cells. J Cell Biol. 193:97-108.
Schoenfelder S, Sexton T, Chakalova L, Cope NF, Horton A, Andrews S, Kurukuti S, Mitchell JA, Umlauf D, Dimitrova DS, Eskiw CH, Luo Y, Wei CL, Ruan Y, Bieker JJ, and Fraser P. (2010) Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat. Genet. 42:53-61.
Hadjur S, Williams LM, Ryan NK, Cobb BS, Sexton T, Fraser P, Fisher AG, Merkenschlager M. (2009) Cohesins form chromosomal cis-interactions at the developmentally regulated IFNG locus. Nature 460,410-413.
Redrup L, Branco MR, Perdeaux ER, Krueger C, Lewis A, Santos F, Nagano T, Cobb BS, Fraser P, Reik W (2009) The long noncoding RNA Kcnq1ot1 organises a lineage-specific nuclear domain for epigenetic gene silencing. Development 136,525-530.
Nagano T, Mitchell JA, Sanz LA, Pauler FM, Ferguson-Smith AC, Feil R and Fraser P (2008) The Air non-coding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322,1717-1720.
Schoenfelder S and Fraser P (2008) Interchromosomal huddle kickstarts antiviral defense. Cell 134,14-16.
Mitchell JA, and Fraser P (2008) Transcription factories are nuclear sub-compartments that remain in the absence of transcription. Genes Dev. 22,20-25.
Sexton, T, Schober, H, Fraser, P, Gasser, S. (2007) Gene regulation through nuclear organization. Nat Struct Mol Biol. 14, 1049-1055.
Osborne, C., Chakalova, L., Mitchell, J., Horton, A., Wood, A., Bolland, D., Corcoran, A. and Fraser, P. (2007) Myc dynamically and preferentially relocates to a transcription factory occupied by Igh. PLoS Biology 5:1763-1772.
Fraser, P. and Bickmore W. (2007) Nuclear organisation of the genome and the potential for gene regulation. Nature, 447, 413-417.
Fraser, P. (2006) Transcriptional control thrown for a loop. Curr Opin Genet and Dev. 16,490-495.
Chakalova L, Debrand E, Mitchell JA, Osborne CS, Fraser P. (2005) Replication and transcription: shaping the landscape of the genome. Nat Rev Genet. 6, 669-677.
West A.G. and Fraser P. (2005) Remote control of gene transcription. Hum. Mol. Genet. 14:R101-111.
Osborne, C.S., Chakalova, L., Brown, K.E., Carter, D., Horton, A., Debrand, E., Goyenechea, B., Mitchell, J.A., Lopes, S., Reik, W. and Fraser, P., (2004) Active genes dynamically co-localize to shared sites of ongoing transcription. Nat. Genet. 36, 1065-1071.
Hacein-Bey-Abina S., Von Kalle C., Schmidt M., McCormack MP., Wulffraat N., Leboulch P., Lim A., Osborne C., Pawliuk R., Morillon E., Sorensen R., Forster A., Fraser P., Cohen J., de Saint Basile G., Alexander I., Munich, Frebourg T., Romana S., Radford J., Gross F., Valensi F., Delabesse E., MacIntyre E., Sigaux F., Soulier J., Leiva L.E., Wissler M., Rabbitts T.H., Le Deist F., Fischer A., Cavazzana-Calvo M. (2003) LMO-2 associated clonal T cell proliferations in two patients after gene therapy for SCID-X1. Science, 302:415-419.
Carter, D., Chakalova, L., Osborne, C.S., Dai, Y-F, Fraser, P. (2002) Long-range chromatin regulatory interactions in vivo. Nat. Genet., 32,623-626.
Milot, E., Strouboulis, J., Trimborn, T., Wijgerde, M., de Boer, E., Langeveld, A., Tan-Un, K., Vergeer, W., Yannoutsos, N., Grosveld, F., Fraser, P. (1996) Heterochromatin effects on the frequency and duration of LCR-mediated gene transcription. Cell 87,105-114.
Wijgerde, M., F. Grosveld and P. Fraser. (1995) Transcription complex stability and chromatin dynamics in vivo. Nature 377; 209-213.
