Our research makes use of maize (corn, Zea mays L.) as a model genetic system to explore hereditary mechanisms and genetic regulation at the molecular, genomic, and cellular levels. We maintain a long-standing focus on the dynamics of telomeres and nuclear envelope interactions in chromosome behavior, with more recent exploration of plant and human chromatin structure, advancing strategies for epigenomic cartography to investigate the genome's response to environmental stimuli. Our work extends to investigating G-quadruplex DNA in stress response and DNA replication in meristem cells, utilizing a diverse array of experimental techniques from traditional genetics to advanced 3D microscopy and bioinformatics.
For links to our research articles, visit the Bass
Lab Publications page.
Chromatin Structure Profiling with MNase:
The DNS-seq and MOA-seq assays, conceived in collaboration with Dr. Jonathan Dennis and others at FSU, create new ways to explore and understand plant and animal chromatin.
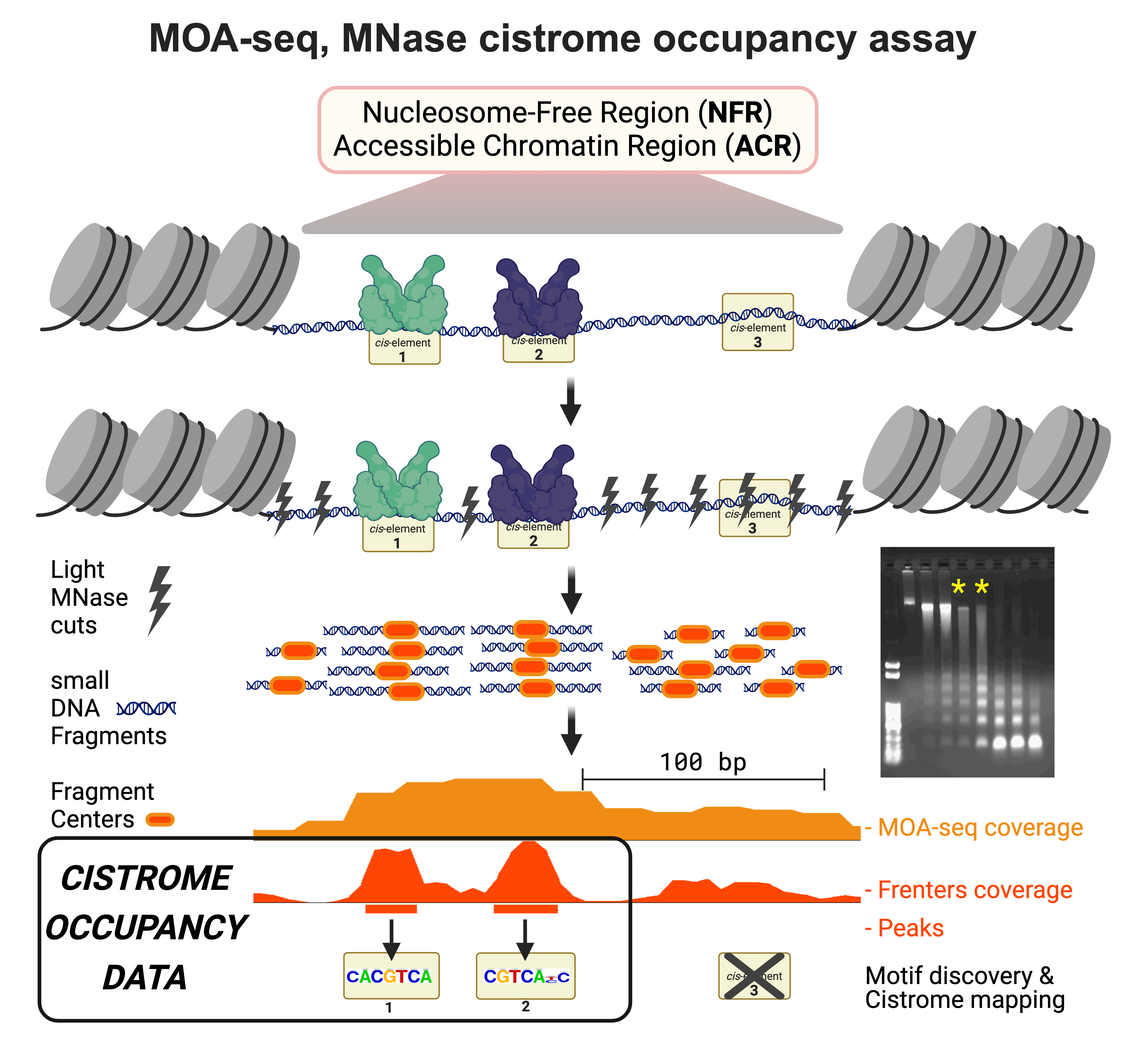
Our techniques repurpose the enzyme MNase for chromatin sensitivity profiling (using DNS-seq) and transcription factor occupancy mapping (using MOA-seq).
The NSF Plant Genome Research Program has generously supported our efforts to discover, develop, and train others to use these approaches.
These methodologies are increasingly being applied to understand the how chromatin structure and genome response are coordinated.
The DNS-seq technique utilizes light and heavy MNase digests to map nuclease hypersensitive regions.
More recently, our MOA-seq approach reported in 2019, has been shown to uncover high-resolution TF-binding at cis-regulatory element sites and motifs.
These techniques contribute to genomic innovation by pinpoint mapping the protein-DNA switches in their native locations.
This knowledge helps us understand gene regulatory networks at occupied cistrome level.
By mapping causal factors regulating heritable phenotypic variance in complex traits, this work enables experimental and applied precision medicine and agriculture.
Genome Replication:
In collaboration with colleagues at NC State University, our lab has pioneered detailed quantitative analyses of DNA replication during the S-phase in maize. These groundbreaking studies have shed light on the existence of distinct chromatin compartments within the nucleoplasm, characterized by their condensation state and replication timing. Supported by the NSF Plant Genome Research Program, our ongoing research employs cutting-edge 3D microscopy, chromatin profiling, and Hi-C techniques to test our models in maize and sorghum, confirming the predictive value of open chromatin for early-S replication.
Notable publications in this area include insights into Arabidopsis DNA replication initiation in open chromatin (Wheeler et al., 2020), reflections on plant DNA replication (Savadel & Bass, 2017), and defining DNA replication patterns in developing maize (Bass et al., 2015; 2014).
Meiotic Nuclear Envelope Telomere Bouquet:
This research was initiated during HWB's postdoctoral research.
Since then investigations have continued to focus on the role of the telomere bouquet in meiosis. Research led by postdocs, graduate and undergraduate students at FSU has led to
the identification, cloning, and characterization of genes encoding maize telomeric proteins,
the
characterization and cloning of meiotic mutants with meiotic telomere-misplacement phenotypes,
QTL mapping of genetic factors that control variation in telomere length,
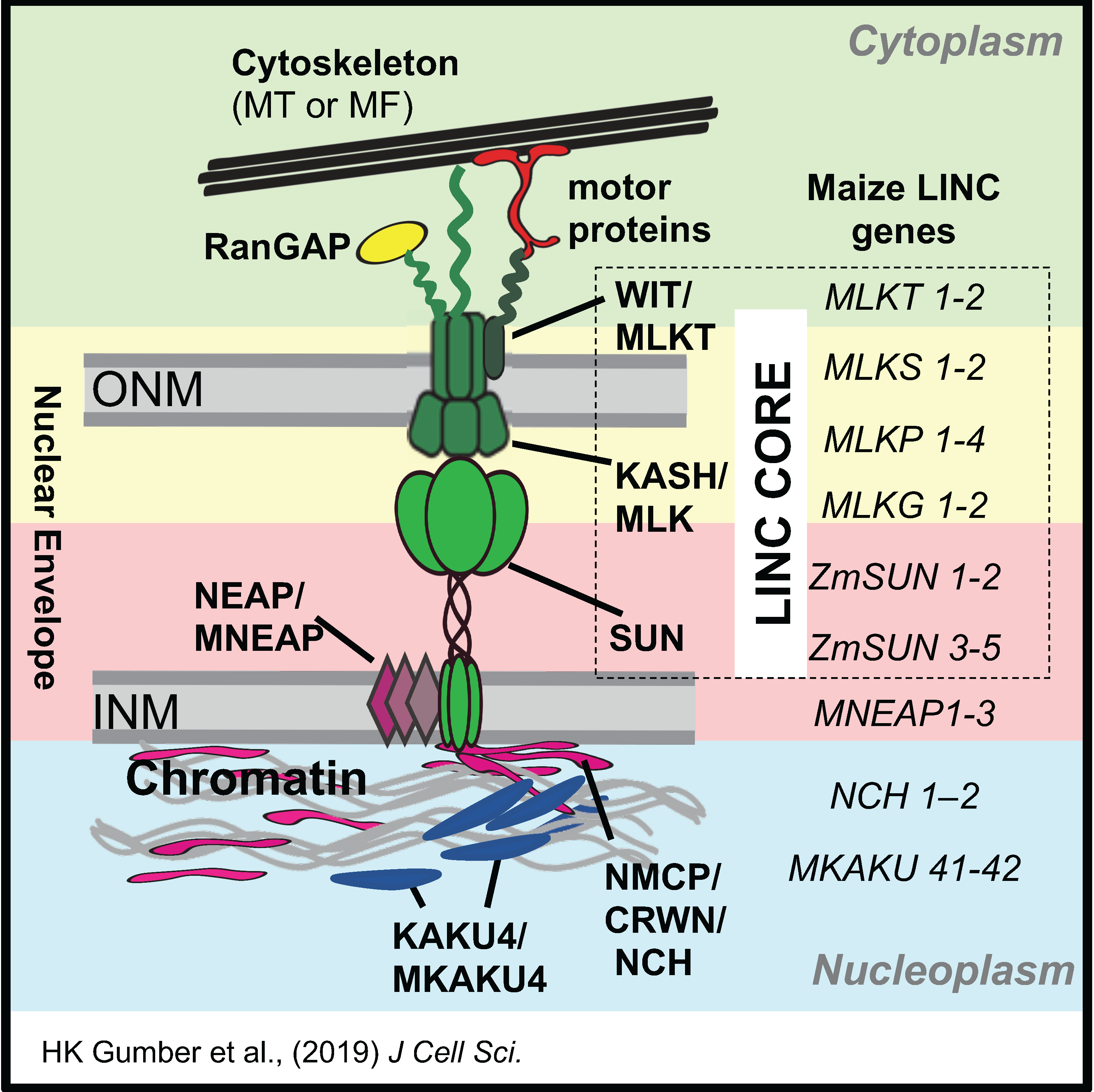
and the analysis of the meiotic nuclear envelope, SUN domain proteins, and the LINC complex
in coordinating the cytoplasm-nucleoplasm interactions that control many aspects of chromosome behavior in meiotic prophase.
New Research Areas
G4 DNA and hypoxia:
Intrigued by the interaction between the maize Shrunken1 (Sh1) gene promoter and ZmIBP1—a telomere-binding protein from the TRFL family—and the presence of a potential G-quadruplex (G4) within its promoter, we did a genomic survey to identify G4 DNA motifs in maize (Andorf et al., 2014).
This venture was a collaborative effort with bioinformaticians, hypoxia specialists, and experts in sugar metabolism from FSU, Iowa State University, and the University of Florida.
We identified a predicted G4 hotspot in thousands of maize genes that tended to be enriched for genes associated with energy stress signaling pathways, including those for hypoxia, sugar stress.
Our results indicate that G4 DNA could influence regulatory processes at the onset of transcription.
In searching for maize G4-binding proteins, we discovered ZmNDPK1, the first report of a plant G4-DNA binding protein (Kopylov et al., 2015).
The plant ZmNTKP1 gene is homologous to human NM23-H2, an NDPK protein associated with G4-mediated transcription at the oncogene MYC.
Our research is now advancing into the biomedical field, through a collaboration with Dr. M Gillespie and colleagues at USA-COM.
We are investigating the possible causal relationships between hypoxia and various pan-genomic responses, including 8-oxoG base damage signaling, G4s, transcriptome remodeling, and chromatin structure.
Prior Research Projects
The Cytogenetic Map of Maize project employs an innovative technique based on transgenomic FISH mapping with maize marker-selected sorghum BAC probes on maize meiotic fibers that are visually isolated by chromosome painting in an oat genome background.
The Chromatin Structure and Genome Response in Maize project
combines genome-wide microarray analysis with 3D microscopy using novel assays
to characterize the nuclesomal and chromatin accessibility landscapes in the
maize genome in response to genetic or experimental manipulations.
This
team project includes investigators from FSU (JH Dennis, JM McGinnis) and Florida A & M University (O Onokpise).
More information on the research projects can be found on the Research page.
3-D Deconvolution Images and Movies:
- Annotated DAPI images of meiotic prophase, (leptotene, zygotene, pachytene, diplotene, diakinesis)
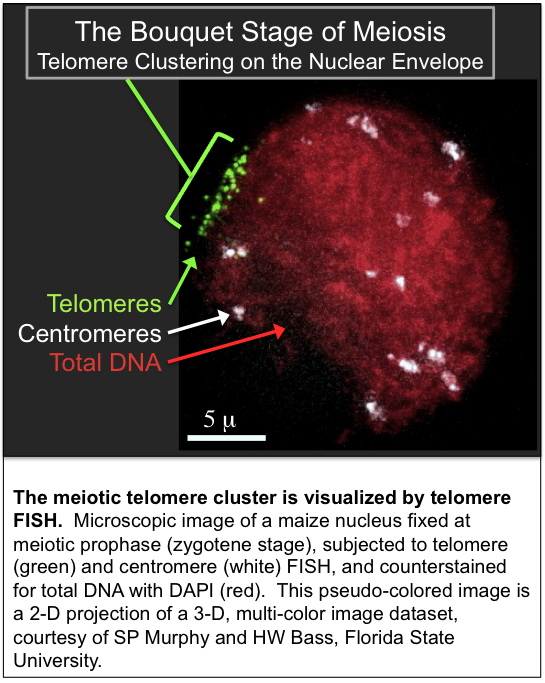
- Telomere FISH at the bouquet stage of meiosis
- Whole chromosome painting analysis of meiotic chromosome pairing in Oat-Maize9
- 3D Images of desynaptic (dy, dsy1) mutants.
- TGMV geminvirus intranuclear localization.
|
|

|