Research in the Hughes Lab
Overview:
Our goal is to understand how natural selection mediated by the physical, biological, and social environment, interacts with other evolutionary processes to maintain diversity in ecologically important traits. We want to know how much genetic diversity is adaptive and how much is non-adaptive, and we are interested in the consequences of both kinds of variation for individuals, populations, and communities.
We work mainly with natural populations of poeciliid (livebearing) fish and fruit flies, using techniques such as field studies, behavioral experiments, and genetic and genomic analysis. Members of the lab are studying the interaction between genetics and social environment in the development of alternative life history traits and reproducitve behavior, the genetic and genomic consequences of sexual selection and mate choice; genetic and social modifiers of aggression and dominance, and the genetic and evolutionary determinants of longevity.
Current projects:
Social interactions and the maintenance of alternative male life histories in sailfin mollies
 Phenotypic variation is a prerequisite for natural selection, but when selection favors a particular phenotype, variation can be quickly eroded. Herein lies a central paradox of evolutionary biology: how is variation maintained in the face of selection on adaptive traits? Alternative life histories are a particularly striking example of variation in adaptive traits. Social interactions are implicated as a crucial factor in the maintenance of phenotypic variation in some species because the fitness consequences of alternative life history phenotypes are typically mediated by social interactions, and development of alternative phenotypes can be regulated by social factors. This project is testing the hypothesis that variation in the social environment is critical to maintaining alternative life history strategies in sailfin mollies, a species that demonstrates dramatic variation in male size and maturation rate within and among populations. We are also assessing whether LH variation is due in part to predictive adaptive response or socially-cued anticipatory plasticity; i.e. that juveniles use social cues to alter their development in order to match their phenotype at maturation to the social environment they will encounter as adults. We are using experiments, field studies, and theoretical models to test these hypotheses, and this project is a collaborative effort between multiple research groups at FSU.
Phenotypic variation is a prerequisite for natural selection, but when selection favors a particular phenotype, variation can be quickly eroded. Herein lies a central paradox of evolutionary biology: how is variation maintained in the face of selection on adaptive traits? Alternative life histories are a particularly striking example of variation in adaptive traits. Social interactions are implicated as a crucial factor in the maintenance of phenotypic variation in some species because the fitness consequences of alternative life history phenotypes are typically mediated by social interactions, and development of alternative phenotypes can be regulated by social factors. This project is testing the hypothesis that variation in the social environment is critical to maintaining alternative life history strategies in sailfin mollies, a species that demonstrates dramatic variation in male size and maturation rate within and among populations. We are also assessing whether LH variation is due in part to predictive adaptive response or socially-cued anticipatory plasticity; i.e. that juveniles use social cues to alter their development in order to match their phenotype at maturation to the social environment they will encounter as adults. We are using experiments, field studies, and theoretical models to test these hypotheses, and this project is a collaborative effort between multiple research groups at FSU.
Behavioral and genomic mechanisms of frequency-dependent selection in guppies
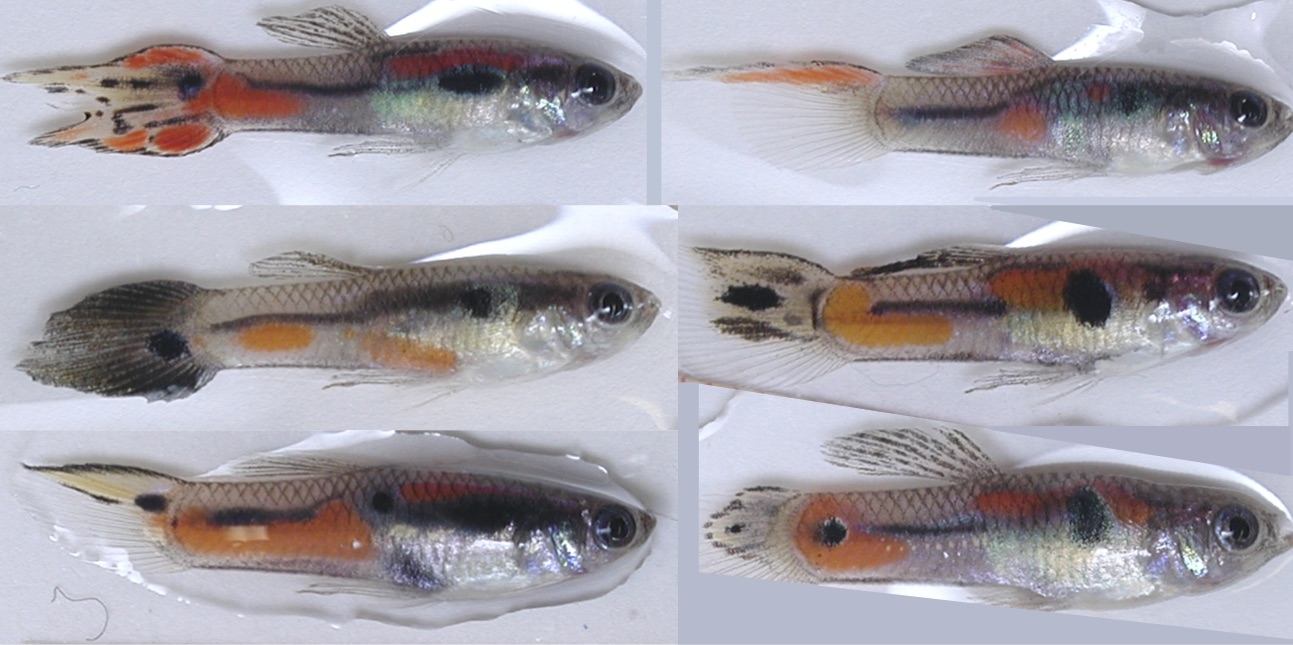 In the genomic era, it has become increasingly obvious that most traits that are relevant to the health, longevity, and fitness of organisms are influenced by genetic variation. Nevertheless, the processes responsible for maintaining genetic variation are generally not known. One reason is that direct experimental investigations are impossible in humans and difficult and expensive in most model organisms. More tractable species that can be studied in both natural and laboratory settings are needed to address this question. Wild guppies exhibit one of the most striking examples of genetic varation known (color pattern variation, see Figure at left for examples of different color patterns found within a single population). This variation is thought to be maintained because individuals with rare color patterns have higher fitness in nature than individuals with common patterns. This project is elucidating the ecological, behavioral and genetic mechanisms underlying these patterns. Specifically, we are investigating social interactions between individuals and ecological interactions between guppies and their predators that generate a selective advantage for rare and novel genotypes. This work will provide a clear picture of how ecological, social and molecular processes contribute to genetic variation.
In the genomic era, it has become increasingly obvious that most traits that are relevant to the health, longevity, and fitness of organisms are influenced by genetic variation. Nevertheless, the processes responsible for maintaining genetic variation are generally not known. One reason is that direct experimental investigations are impossible in humans and difficult and expensive in most model organisms. More tractable species that can be studied in both natural and laboratory settings are needed to address this question. Wild guppies exhibit one of the most striking examples of genetic varation known (color pattern variation, see Figure at left for examples of different color patterns found within a single population). This variation is thought to be maintained because individuals with rare color patterns have higher fitness in nature than individuals with common patterns. This project is elucidating the ecological, behavioral and genetic mechanisms underlying these patterns. Specifically, we are investigating social interactions between individuals and ecological interactions between guppies and their predators that generate a selective advantage for rare and novel genotypes. This work will provide a clear picture of how ecological, social and molecular processes contribute to genetic variation.
Social interactions and the maintenance of genetic polymorphism in the eastern mosquitofish, Gambusia holbrooki
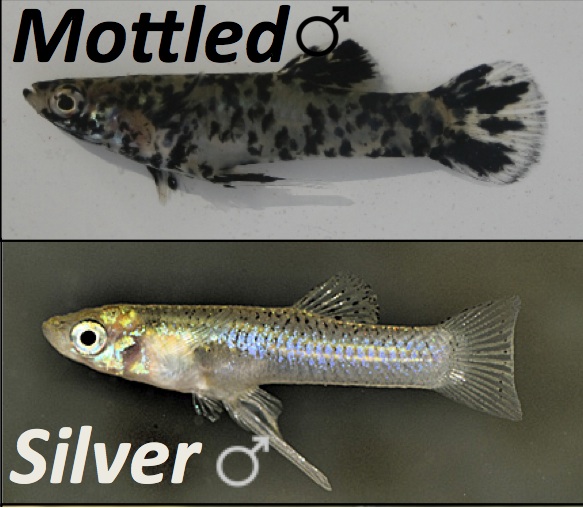 For decades biologists have struggled to determine if individual behavior is influenced more by nature (genes) or nurture (social environment). However, emerging research suggests that nature and nurture are complex phenomena that interact at many levels. Individuals that differe genetically might respond differently to the same environmental or social cues, and indivuals might direct different behaviors toward individuals with different genotypes. Morevover, these "indirect genetic effects" can feed back one each other to influence individual fitness and population dynamics. One way to make this problem more tractable is to use a species in which individuals differ in behavior because of a single gene. We are using mosquitofish that exhibit two different genetically determined color patterns within natural populations (see Figure at right), that also differ in aggression and other social beahviors.
For decades biologists have struggled to determine if individual behavior is influenced more by nature (genes) or nurture (social environment). However, emerging research suggests that nature and nurture are complex phenomena that interact at many levels. Individuals that differe genetically might respond differently to the same environmental or social cues, and indivuals might direct different behaviors toward individuals with different genotypes. Morevover, these "indirect genetic effects" can feed back one each other to influence individual fitness and population dynamics. One way to make this problem more tractable is to use a species in which individuals differ in behavior because of a single gene. We are using mosquitofish that exhibit two different genetically determined color patterns within natural populations (see Figure at right), that also differ in aggression and other social beahviors.
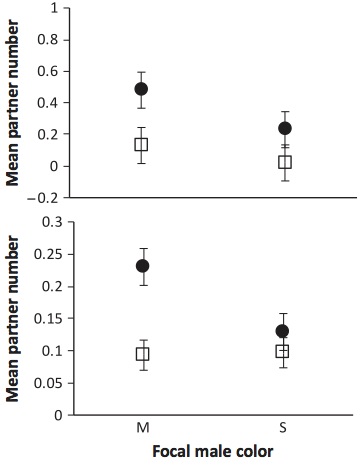
These experiments will determine how group composition affects the behavior of juvenile and adult fish. For example, how does living in a high aggression group affect the health and behavior of a juvenile fish, and does the effect of the social environment depend on the juveniles own genetic makeup? This research will determine if these social effects are responsible for the maintenance of these two genetically distinct types within the same local populations. That is, do social interactions help to maintain genetic diversity? The plot to the left shows that mottled males associate with more females (filled symbols) that do silver males, but both types of males associated equally frequently with other males. This pattern holds both in the field (top panel) and in the lab (bottom panel)
Evolutionary lability and adaptive plasticity in mechanisms of behavior
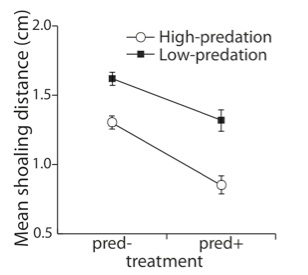 Genes shape how individuals respond to their environment, and this environmental sensitivity influences behavior, life history, and fitness. This project seeks to understand how genes and developmental conditions together influence the brain, and how that interaction alters social behavior. We take advantage of extensive information on genetic and environmental influences on the behavior of guppies, small fish that have evolved numerous behavioral responses to predators. Guppies that experience high and low levels of predation in the wild are being raised in laboratory conditions with and without predator exposure. Genetic and molecular experiments will link the patterns of gene expression in different brain regions to neural activity patterns and to the resulting social behaviors. Results will demonstrate whether similar behavioral traits (e.g., increased sociality in fish from high-predation locales and in fish exposed to predators) rely on the same genomic and brain activity patterns, or if they can emerge from a variety of neural mechanisms. These findings will also reveal how sensitivity to the environment shapes the evolution of behavior. Our novel experimental approach will provide a model for other researchers seeking to understand the impacts of gene expression differences on behavior.
Genes shape how individuals respond to their environment, and this environmental sensitivity influences behavior, life history, and fitness. This project seeks to understand how genes and developmental conditions together influence the brain, and how that interaction alters social behavior. We take advantage of extensive information on genetic and environmental influences on the behavior of guppies, small fish that have evolved numerous behavioral responses to predators. Guppies that experience high and low levels of predation in the wild are being raised in laboratory conditions with and without predator exposure. Genetic and molecular experiments will link the patterns of gene expression in different brain regions to neural activity patterns and to the resulting social behaviors. Results will demonstrate whether similar behavioral traits (e.g., increased sociality in fish from high-predation locales and in fish exposed to predators) rely on the same genomic and brain activity patterns, or if they can emerge from a variety of neural mechanisms. These findings will also reveal how sensitivity to the environment shapes the evolution of behavior. Our novel experimental approach will provide a model for other researchers seeking to understand the impacts of gene expression differences on behavior.






